Yifan Zhuo – Life Science
This is a pre-print of a manuscript currently under peer-review.
Abstract
CRISPR genetic editing technique is more prevalently used in healthcare and clinical settings in these days. This study aims to discover the viability of genetically modified Escherichia coli (E. coli) from a miniPCR bioTM CRISPR/Cas Gene Knockout Kit in acidic environments, like that found in the human stomach. It was observed that the genetically modified E. coli grew successfully in the presence of acid. The unexpected color patterns of blue dots and rings in the white bacteria colonies sparked a second experiment analyzing the interaction between acid and X-gal staining. However, the blue dots and rings were not observed in the second experiment possibly due to the halted bacteria growth and reduced volume of acid and ChIX used in the second experiment. The interaction between acid and X-gal staining has not been studied in existing literature. Therefore, it is suggested that this study should be repeated in the future, where a bacteria strain whose growth is not inhibited by strong acid is used and more bacteria colonies are successfully grown.
Introduction
In recent years of rapid advancements in the field of biotechnology, one of the most discussed gene technologies has been clustered regularly interspaced short palindromic repeats, which can be abbreviated as CRISPR (Liu et al., 2021). More specifically, CRISPR-Cas9 is by far the most extensively used gene-editing nuclease. This approach uses specially designed guide RNA and Cas protein to cut or replace specific genes in the organisms. CRISPR-Cas9 has great promise for different disease treatments, including neurodegenerative diseases, cancers, and so on (Alagoz & Kherad, 2020). In these treatments, editing host cells in vitro and transplanting them back into the host can enable edited cells to possess high specificity and compatibility with host organisms (Faintuch & Faintuch, 2019).
A potential example of the application of CRISPR-Cas9 is microbiota transplantation (Ramachandran & Bikard, 2019). So far, microbiota transplantation is considered safe and well-tolerated in high-risk patients. Studies indicate that microbiota transplantation is a promising treatment for gastric diseases. In one paper, it is mentioned that CRISPR interference (a technique used to repress specific gene expressions) was used on microbiome-associated bacteria, including Lactococcus lactis and Bacteroides thetaiotaomicron, to alter the bacterial metabolic capacity and its resistance to antimicrobial peptide (Park & Seo, 2021). However, there have been concerns regarding danger of transmitting infectious pathogens from donor to patient. With the rapid development and manipulation of gene editing technology, it is possible to implant genetically edited bacteria extracted from patients themselves to mitigate risks of acquiring unknown pathogens (Ramachandran & Bikard, 2019). This may minimize the negative impacts of non-host microbiota transplantation. Nevertheless, there have not been studies addressing the viability of gene edited bacteria in low pH environment like the human stomach, which will be the goal of this study. Escherichia coli (E. coli) has long been a model system in labs, which reactions to stimuli in lab can be expected to occur similarly in the human microbiota (Klymkowsky & Cooper, 2016). In this study, E. coli will be grown in the presence of hydrochloric acid (HCl) solutions with pH 1.5 and 2.0, corresponding to the upper and lower boundaries of human stomach acidity range (Fujimori, 2020). The E. coli used in the experiment will come from miniPCR bioTM CRISPR/Cas Gene Knockout Kit and will contain pLacZ plasmids that carries the lacZ gene. The lacZ gene encodes β-galactosidase enzyme, which catalyzes the breakdown of the sugar lactose. The β-galactosidase enzyme also catalyzes the breakdown of the chemical X-gal, which will show a blue color when broken down in this experiment. If the lacZ gene is knocked out using specific guide RNA for lacZ and cas9 gene, the X-gal will not be broken down, and resulting bacteria colonies will show a white color instead. This study will compare genetically modified E. coli with the lacZ gene knocked out, with control E. coli to find out the viability of genetically modified E. coli in the presence of strong acid. This study will also reveal whether the presence of acid will affect the inability of genetically modified E. coli to break down X-gal, which acts as a proxy for lactose. This will be shown by the color of the resulting bacteria colonies. If the genetically modified bacteria show a white color, the bacteria still cannot break down X-gal, and therefore lactose, as expected.
Materials and Methods
A miniPCR bioTM CRISPR/Cas Gene Knockout Kit was used to conduct the contrast experiment between genetically modified and control E. coli. All Gene Knockout Kit related methods were used according to miniPCR kit’s instruction (miniPCR Lab Knockout Instructor Guide, 2023).
Preparing Agar Plates with X-gal
Inside the miniPCR bioTM CRISPR/Cas Gene Knockout Kit, a vial labelled ChIX was found and thawed in an incubator set at 37°C. ChIX mix included the X-gal necessary for testing the presence of lacZ gene in E. coli and therefore the ability of E. coli to break down lactose. A 250 ml graduated cylinder was used to measure out 190 ml of distilled water, which was added to a 500mL heat-proof bottle with 7.5 g of Luria broth (LB) agar powder. After fully dissolving the powder by swirling, the bottle was loosely capped and put into a microwave. The bottle was checked every 30 seconds and carefully swirled until the solution was visibly homogenous and contained some bubbles. The LB agar solution was cooled for about 4 minutes until the bottle could be touched with bare hands. 475 μl of ChIX was added to the LB agar solution using a p1000 pipette and mixed by swirling. Eight Merlan 15 ml compartmented sterile Petri dishes were laid out, with each dish containing three compartments. Finally, the agar solution was poured into each compartment of the Petri dishes to a thickness of 3 mm. While pouring agar, two candles were placed near the plates to prevent possible contamination. The lids were immediately put on the Petri dishes, and LB agar was allowed to solidify at room temperature.
Preparing Acidic Solutions (Upper and Lower Boundaries of Human Acid Range: pH 1.5 and 2.0)
The Westlab 3%, 1M, HCl was used to produce acidic solutions of pH 1.5 and 2.0. In each of the two 500 ml beakers, 200 ml of distilled water was poured. The two beakers were labelled: one as “pH 1.5” and the other as “2.0”. Approximately 50 ml of the HCl was poured into a 100 ml beaker. 1000 μl of HCl was added each time into the “pH 1.5” beaker using a p1000 pipette until the mixture reached pH 1.5, the mixture was swirled thoroughly using a plastic pipette (acting as a stirring rod), and an electronic pH meter (Dr. Meter) was used to measure the acidity of the mixture after each addition. The pipette tips and plastic pipettes were discarded, and the pH meter was cleaned using water between samples to avoid inaccurate measurements. The pH 2.0 solution was made using the same procedure. The two beakers were covered using plastic wrap to prevent contamination.
Preparing Bacteria for Transformation Experiment
Three days before the bacteria transformation lab, 900 μl of LB+ ampicillin liquid media were added to the vial of lyophilized E. coli (included in miniPCR kit) using a p1000 pipette after removing the rubber stopper on the vial. After reinserting the rubber stopper, the vial was inverted several times to mix. The 50 μl of rehydrated bacteria were aliquoted into eight sterile 1.7 ml microtubes using a p100 pipette. The microtubes were incubated at room temperature for 48 hours.
One day before the bacteria transformation lab, 25 μl of sterile CaCl2 was added to each tube of bacteria using a p100 pipette. Pipette tips were changed between samples to prevent contamination. The tubes were placed in refrigerator and chilled for 24 hours.
lacZ Gene Knockout in E. coli
After thawing the eight 1.7 ml tubes that contained bacteria with hands, the tubes were labelled: four tubes as “Ctrl rxn” (control reaction) and the other tubes as “KO rxn” (for knockout reaction). The tubes were placed on ice. 25 μl of pCtrl DNA (plasmid containing Cas9 gene and random guide RNA, which cannot successfully knock out lacZ gene) was added to each tube labelled “Ctrl rxn” using a p100 pipette, and 25 μl of pKO DNA (plasmid containing Cas9 gene and guide RNA for lacZ, which targets the lacZ gene for knockout) was added to each tube labelled “KO rxn” using a p100 pipette. Pipette tips were changed between samples to prevent contamination. The tubes were securely closed, flicked to mix the solution inside, and tapped on the bench to collect liquid in the tube bottom. The tubes were then placed on ice so that they had sufficient contact with ice for 30 minutes. After that, the tubes were placed in a water bath set at 42°C for 90 seconds. Then, the tubes were incubated on ice again for 2 minutes. 50 μl of recovery media (which is formulated to ensure high bacteria transformation) were added to each of the tube using a p100 pipette, and pipette tips were changed between samples to prevent contamination. The tubes were securely closed, flicked to mix the solution inside, and tapped to collect liquid in the tube bottom. Then, the tubes were put in the incubator set at 37°C for 20 minutes.
Plate Transformed and Control Bacteria onto X-gal Agar Plates
The eight agar plates containing X-gal were labelled: four plates as “Ctrl rxn” and the other four plates as “KO rxn”. 33 μl of solution in “Ctrl rxn” tube was added to each compartment on the “Ctrl rxn” plate, and 33 μl of solution in “KO rxn” tube was added to each compartment on the “KO rxn” plate using a p100 pipette. After adding the solution, the mixture was spread evenly across the surface of the agar using a glass spreader sterilized on flame between samples. The lids were immediately put on Petri dishes.
Add Acids to Appropriate Plate Compartments
After the plates sat for about 5 minutes following plating, appropriate acids were added into corresponding plate compartments. The compartmentalized plates used in this study had pre-labelled numbers for each compartment (I, II, III). For all eight plates, 300 μl of pH 1.5 acidic solution was added to compartment I, and 300 μl of pH 2.0 acidic solution was added to compartment II. No acid was added to compartment III, which served as control groups. The plates with added acids were carefully swirled to maximize area covered by acids. Finally, all plates were incubated at 25°C overnight.
Transfer Grown Control and KO Bacteria
This step was completed because limited bacteria colonies were grown on the first incubated plates. Two sterile, single-compartment Petri plates were labelled as “KO no acid” and “Ctrl no acid”. Bacteria in compartment III (no acid) of the grown KO plate were transferred and streaked onto the new “KO no acid” plate, and bacteria in compartment III (no acid) of the grown Ctrl plate were transferred and streaked onto the new “Ctrl no acid” plate using a sterile pipette tip. Pipette tips were changed between samples to prevent contamination. During the whole process, two candles were placed near the plates to prevent possible contamination. Finally, all plates were incubated at 25°C overnight.
Interaction between X-gal and Acid
After observing the unusual colonies with blue and white color patterns shown by the transformation experiment, a second part of the research was conducted to investigate the interaction between X-gal and HCl. A 500 ml graduated cylinder was used to measure out 150ml agar solution. After the LB agar solution was cooled enough to be touched with bare hands, 120 μl of ChIX was added to the solution using p200 pipette and swirled to mix. Six sterile, single-compartment Petri dishes were labelled: KO no acid, KO pH 1.5, KO pH 2.0, Ctrl no acid, Ctrl pH 1.5, and Ctrl pH 2.0. The agar solution was poured into six dishes. Bacteria from the plate labelled “Ctrl no acid” grown in the first part of the experiment were transferred and streaked onto the three new Control plates, and bacteria from the plate labelled “KO no acid” were transferred and streaked onto the three new KO plates using a sterile pipet tip. After remeasuring the pH of pre-prepared acidic solutions, 300 μl of pH 1.5 solution was added to the pH 1.5 plates, and 300 μl of pH 2.0 solution was added to the pH 2.0 plates using a p1000 pipette. The plates with added acids were carefully swirled to maximize area covered by acids. Finally, all plates were incubated at 25°C for one week when the first observation of results took place.
Results
Genetically Modified Bacteria’s Survival in Acid
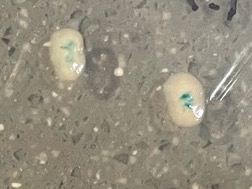
Overall, the KO plates produced white colonies, and Ctrl plates produced white colonies, as expected. However, as shown in Table 1, there were four white colonies on KO plates that showed tiny blue dots in the middle of the colonies. An example of the unusual color patterns is shown in Figure 1, where two white colonies are grown with blue dots in the middle.
Table 1. First Week after Bacteria Transformation Experiment
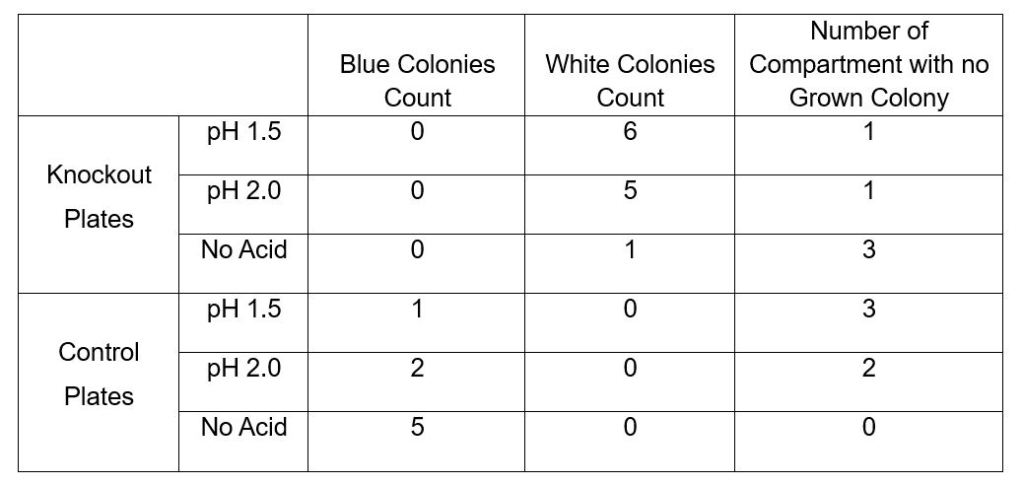
Figure 1. White colonies with blue dots (grown in presence of pH 1.5 acid on a knockout plate) after one week.
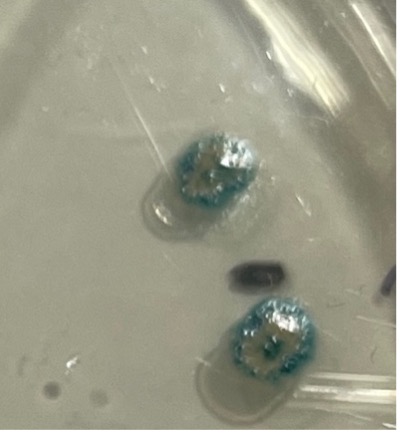
Two weeks after the bacteria transformation experiment, the total number of colonies grown for each compartment in plates did not change, but the colonies all grew larger in size. The tiny blue dots seen in the middle of some white colonies got larger, more blue dots emerged around the dots in the colonies, and blue rings formed at the colonies’ edges, as shown in Figure 2.
Figure 2. The same two colonies seen in Figure 1 had more blue dots emerge in the middle, and blue rings formed at the edges, after the second week.
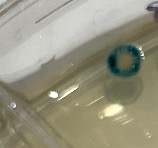
Other white colonies that did not show tiny blue dots showed significant blue rings at the colonies’ edges, as shown in Figure 3. The blue colonies did not show an obvious mixing color pattern.
Figure 3. A white colony (grown in the presence of pH 2.0 acid on a knockout plate) that did not have blue dots in the first week showed dark blue rings at the edges after the second week.
Interaction between Acid and X-gal
Due to the aforementioned unusual color patterns of grown colonies, a second experiment was conducted to investigate the interaction between HCl acid and X-gal. As shown in Figure 4 and 5, first and second week after the acid X-gal interaction experiment, regardless of the presence of acid, all three plates labelled as KO contained uniform white bacteria colonies, while all three plates labelled as Ctrl contained uniform blue bacteria colonies.
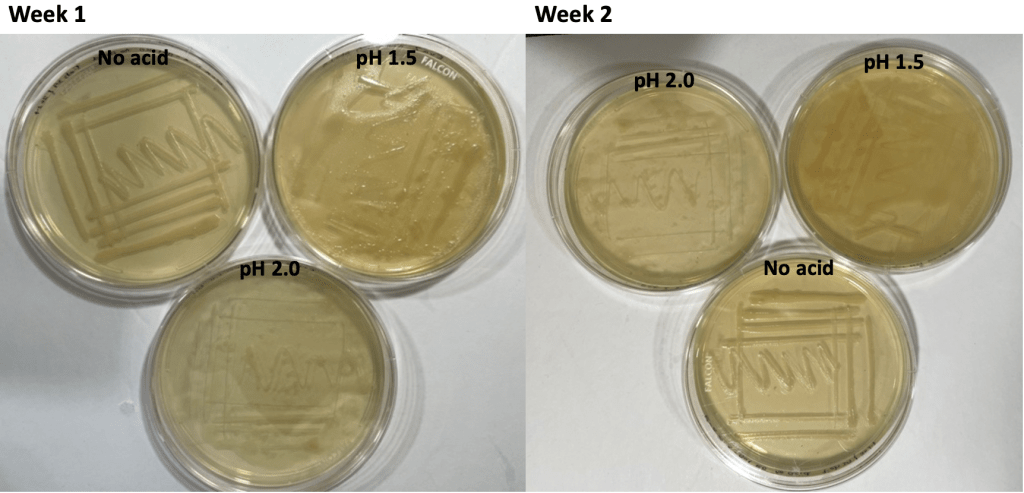
Figure 4. Knockout bacteria colonies

Figure 5. Control bacteria colonies
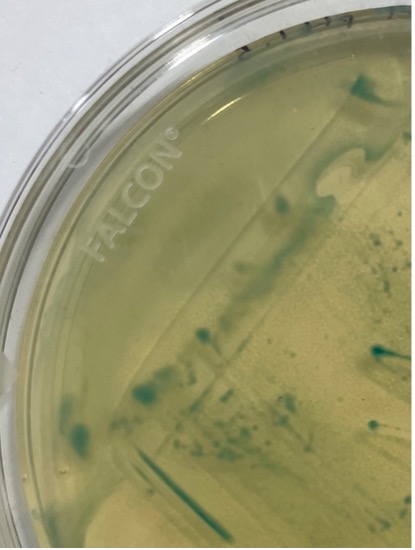
The only difference of bacteria growing pattern between the six plates was that the bacteria colonies in acids grew all over the plates, in a hazy pattern, rather than growing in a distinct streaking pattern, due to the presence of acid (Figure 6). However, it is worth noting that the bacteria stopped growing as the bacteria colonies’ size stayed the same after the first week.
Figure 6. A large, light blue section grown at the edge of the plate (knockout plate in presence of pH 1.5 acid) shows a hazy pattern.
Discussion
Genetically Modified Bacteria’s Survival in Acid
The results of the bacterial transformation experiment showed significantly fewer bacteria colonies grown than expected. There were a few limitations and errors that could have caused such results. First, as stated in the experiment instructions, the tubes containing bacteria should be thawed in a water bath at room temperature, and the ChIX LB agar plates should be pre-warmed at 37°C before the experiment. These preparations were not done in this study. Second, when rehydrating the bacteria 3 days before the lab, vials containing bacteria were accidentally spilled. Third, due to time constraint, incubation and icing time was minimized (instead of the suggested 60 minutes incubation, the bacteria were only incubated for 20 minutes). Fourth, each compartment on the three-compartment plate contained 33 μl of bacteria, adding up to 99 μl of bacteria per plate, which was less than the suggested 160 μl of bacteria per plate. Lastly, the transformed bacteria grown on agar plates were incubated at 25°C rather than the suggested 37°C stated on the instruction. All of the errors above might have contributed to the lower number of bacteria colonies grown. The contamination that happened during the experiment likely contributed to the lower number of bacteria colonies grown as well, since the bacteria grown as a result of contamination could compete with target bacteria colonies for limited nutrients in agar (Bacterial Growth n.d.).
Due to the mixed colors of colonies observed after the bacteria transformation experiment, a second part of the experiment was conducted. In KO groups, lacZ was knocked out, and therefore, X-gal was not able to be broken down, showing the color white. It was hypothesized that even though lacZ was not present to break down X-gal, acid might break the X-gal down instead, so a blue color was still shown. In other words, the aim of the second part was to study the possible impact of acid on the breakdown of X-gal.
Interaction between Acid and X-gal
The second part of the experiment showed uniform bacteria colonies’ color, which was expected. It was worth noting that the uniform color pattern on each plate rejected other possible causes of mixing colored colonies on the same plate in the bacterial transformation experiment, including contamination, gene transfer, and mutation. These hypotheses would only be possible if after replating from the first part of the experiment, the colonies again showed mixed color patterns after the first week. However, the hazy bacteria colonies grown on plates, shown in Figure 8, prevented quantitative colony comparisons between plates.
Compared to the bacterial transformation experiment where 300 μl of acid was added to each compartment of the plate and 900 μl of acid was added to each plate, the experiment investigating the interaction between acid and X-gal had 300 μl of acid per plate, which was one-third of the acid amount added in the prior experiment. In the first part of the experiment, far fewer bacteria colonies were successfully grown than expected. It was suspected that the high volume of acid might affect the normal growth of bacteria. Therefore, the change in acid volume in the second part of the experiment was made. 120 μl of ChIX was added to the agar plate rather than 475 μl of ChIX due to the shortage of ChIX solution available. The bacteria appeared to stop growing after one week. It is also worth noting that there was strong evidence of mixed colors after one week which even worsened in the second week in the first experiment. There is no evidence of this in the second experiment. The reduced volume of acid and ChIX and halted bacteria growth might be the reasons why mixed color was not observed in KO.
The findings of the research were consistent with existing literature on the survival of E. coli in acid. In a study, it was found that the acid limit for E. coli growth is pH 4.0 to 4.5, and E. coli can survive in lower pH environment but without growth (Xu et al., 2020). Therefore, the use of pH 1.5 and pH 2.0 HCl solution might inhibit E. coli growth in this study. Even though the overall result shows that genetically modified bacteria can survive successfully in the presence of acid (pH 1.5 and pH 2.0) and still have the inability to take up lactose (as shown by their white color in this kit), the first part of this study should be repeated in the future with more bacteria colonies to show a more accurate simulation experiment and possibly quantify the bacteria colonies grown. Another element to modify in the future study is to use a bacteria strain that might be found in the human stomach and can continue growing in low pH environment. The results from the first part of the experiment pointed at an interesting question of possible interaction between HCl and X-gal, which had not been studied in the literature yet and should be further studied in the future as well.
References
Alagoz, M., & Kherad, N. (2020). Advance genome editing technologies in the treatment of human diseases: CRISPR therapy (Review). International Journal of Molecular Medicine, 46(2), 521–534. https://doi.org/10.3892/ijmm.2020.4609
Bacterial growth—Micro-organisms and their applications – WJEC – GCSE Biology (Single Science) Revision—WJEC. (n.d.). BBC Bitesize. Retrieved April 8, 2024, from https://www.bbc.co.uk/bitesize/guides/z243g82/revision/1
Faintuch, J., & Faintuch, J. J. (2019). Chapter 46 – Precision Medicine: The Microbiome and Metabolome. In J. Faintuch & S. Faintuch (Eds.), Microbiome and Metabolome in Diagnosis, Therapy, and other Strategic Applications (pp. 435–449). Academic Press. https://doi.org/10.1016/B978-0-12-815249-2.00046-4
Fujimori, S. (2020). Gastric acid level of humans must decrease in the future. World Journal of Gastroenterology, 26(43), 6706–6709. https://doi.org/10.3748/wjg.v26.i43.6706
Klymkowsky, M. W., & Cooper, M. M. (2016, June 27). 9.14: E. coli as a model system. Biology LibreTexts. https://bio.libretexts.org/Bookshelves/Cell_and_Molecular_Biology/Book%3A_Biofundamentals_(Klymkowsky_and_Cooper)/09%3A_Genomes_genes_and_regulatory_networks/9.14%3A_E._coli_as_a_model_system
Liu, W., Li, L., Jiang, J., Wu, M., & Lin, P. (2021). Applications and challenges of CRISPR-Cas gene-editing to disease treatment in clinics. Precision Clinical Medicine, 4(3), 179–191. https://doi.org/10.1093/pcmedi/pbab014
miniPCR Lab Knockout Instructor Guide. (2023, February). https://www.minipcr.com/wp-content/uploads/KT180001_miniPCR_Lab_Knockout_instructor_guide_V1.2_010823.pdf
Park, S.-Y., & Seo, G. S. (2021). Fecal Microbiota Transplantation: Is It Safe? Clinical Endoscopy, 54(2), 157–160. https://doi.org/10.5946/ce.2021.072
Ramachandran, G., & Bikard, D. (2019). Editing the microbiome the CRISPR way. Philosophical Transactions of the Royal Society B: Biological Sciences, 374(1772), 20180103. https://doi.org/10.1098/rstb.2018.0103
Xu, Y., Zhao, Z., Tong, W., Ding, Y., Liu, B., Shi, Y., Wang, J., Sun, S., Liu, M., Wang, Y., Qi, Q., Xian, M., & Zhao, G. (2020). An acid-tolerance response system protecting exponentially growing Escherichia coli. Nature Communications, 11(1), 1496. https://doi.org/10.1038/s41467-020-15350-5