Karina Freire Kloecking – Life Science
Abstract
This study aimed to investigate the off-target effects of CRISPR-Cas9 on Escherichia coli (E. coli) DH5α, focusing on the impact of the lacZ gene removal on antibiotic sensitivity and shedding light on CRISPR’s broader implications. Using CRISPR technology and a miniPCR “Knockout! A CRISPR/Cas Gene Targeting Lab” kit, the lacZ gene, known for its metabolic role, was knocked out. Changes in antibiotic susceptibility between the lac-positive and lac-negative bacteria were assessed using the antibiotic disk diffusion method. The results produced minimal observable effects on antibiotic sensitivity in both control and knock-out bacterial strains due to experimental error. While initial consideration of the findings might seem to suggest that the CRISPR off-target effects were minimal, various limitations and errors in experimental design impacted the results, making the data inconclusive. Overall, this study aimed to contribute to the understanding of CRISPR-Cas9 off-target effects and bacterial antibiotic resistance and highlights the need for further study in this area.
Introduction
CRISPR gene editing technology has been groundbreaking in the field of microbiology with its revolutionary precision and speed, but it is subject to numerous discussions and debates over its precision, ethical implications, and more (Lorenzo et al, 2022). CRISPR technology is often described as a genetic pair of scissors, allowing for the precise cut and modification of genes. However, no matter how accurate, there is always the probability of unwanted side effects (Guo et al, 2023). This study will explore the off-target effects of CRISPR-Cas9 technology on E. coli DH5α, specifically when the lacZ gene is targeted. These off-target effects are unintended alterations or mutations that can occur in DNA sequences outside the target site when using this gene editing technology, which could potentially lead to unexpected consequences.
The lacZ gene is best known for its ability to break down lactose (Juers et al, 2012), but this study aims to discover any side effects there may be when it is removed. More specifically, side effects relating to antibiotics. Although the lacZ gene doesn’t have any direct relation to antimicrobial resistance and is not essential to the survival of E. coli (Battad-Bernardo et al, 2004), the off-target effects of CRISPR technology and the removal of this gene may cost the overall fitness of the bacteria (Guo et al, 2023), leading to changes in its susceptibility to antimicrobial substances.
Using the miniPCR “Knockout! A CRISPR/Cas Gene Targeting Lab” to target the lacZ gene and the disk diffusion method to observe changes in antibiotic susceptibility by differences in inhibition zones, this study is about understanding the effects of CRISPR knockout, and seeing how the genetic change might affect the bacteria in unexpected ways. Understanding possible off-target effects is vital when considering the vast potential of CRISPR, and its contributions and potential in all areas of genetic engineering.
Material and Methods
The lacZ gene was knocked out using the miniPCR “Knockout! A CRISPR/Cas Gene Targeting Lab.” The manufacturer’s instructions were followed precisely for this process.
The bacteria from the miniPCR knock-out kit were rehydrated three days in advance for this experiment. To start, 900 μl of LB + ampicillin liquid media was added to the vial of lyophilized bacteria. Once added, the vial was capped with a rubber stopper and inverted several times to mix. The rehydrated bacteria were divided into 16 sterile 1.7 mL microtubes, 50 μl of rehydrated bacteria going into each tube. The samples were then incubated at 25°C for 48-60 hours without being disturbed.
One day before the lab, 25 μl of sterile CaCl2 was added to each tube of bacteria. The tubes were then placed in the refrigerator for 12-24 hours.
ChIX was thawed and set aside, and 190 mL distilled water was added to a 500 mL heat-proof bottle. A bottle was swirled to mix the contents after the addition of 7.5 g LB agar powder. The bottle was microwaved in 30-second increments just until boiling with the bottle loosely capped to allow for steam to vent. The bottle was monitored closely to ensure it didn’t boil over. The solution was carefully swirled using heat-proof mitts. This microwaving process was repeated until LB agar powder was fully dissolved and the solution was transparent. LB agar solution was left to cool until it could be touched with bare hands but not so long that the agar began to sit (about four minutes). After, 475 μl of the fully thawed ChIX was mixed and added to the LB agar solution. The solution was swirled to mix after the addition of the ChIX. Once mixed, the LB agar was poured into eight sterile Petri dishes with a thickness of about three millimeters in each dish. The lids were immediately put back onto the Petri dishes and it was left to solidify at room temperature. Once the agar solidified, the plates were stacked upside down, placed in a plastic bag, and stored in the refrigerator until use.
A water bath was set to 42 °C for heat shock and an incubator was set to 37°C for recovery after transformation and incubating LB agar plates. An ice bath was prepared by filling a large beaker with ice. Before starting the lacZ knock-out procedure, the ChIX LB agar plates were placed in a 37°C incubator.
Tubes containing DNA samples were all thawed by being placed on a rack at room temperature. Four 1.7ml microtubes were labelled and 30 μl of a control plasmid called pCtrl, which contained the cas9 gene and random, insignificant instructions for a guide RNA (gRNA), was dispensed into each. A further four microtubes were labelled and 30 μl of pKO, a plasmid which introduced the cas9 gene and the sequence for a custom gRNA for 20 bases of the lacZ gene, was dispensed into each . Exactly 120 μl SOC recovery media was dispensed into four labelled 1.7 mL microtubes.
Four 1.7 mL tubes containing bacteria were labelled ‘Ctrl rxn’ and another four were labelled ‘KO rxn.’ Once labelled, 25 μl pCtrl DNA was added to each ‘Ctrl rxn’ tube and 25 μl pKO DNA to each ‘KO rxn’ tube, changing pipette tips between samples to prevent contamination. Transformation reaction tubes were securely closed and then flicked a few times to mix the contents. Once mixed, the tubes were tapped on the lab bench to collect liquid at the bottom of the tube. Transformation reaction tubes were then incubated on ice for 30 minutes.
After 30 minutes, the tubes were moved to the water bath for a heat shock, allowing them to sit in the 42°C bath for 90 seconds. The tubes were then once again incubated on ice for two minutes.
After the two-minute incubation, 50 μl recovery media was added into all eight transformation reaction tubes, once again changing pipette tips between samples to prevent contamination. Tubes were securely closed and mixed by being flicked a few times and then tapped on the bench to collect liquid at the bottom of the tube. Following this, the transformation reactions were incubated at 37°C for 20 minutes. While the transformation reaction tubes were being incubated, four LB agar plates were labelled as ‘Ctrl’ rxn and another four were labelled as ‘KO rxn.’
Control transformation reactions were plated using a micropipette to add the entire contents of one ‘Ctrl rxn’ tube (~160 μl) onto one ‘Ctrl rxn’ agar plate and repeating this with each ‘Ctrl rxn’ tube and plate. The mixtures were spread evenly on the surface of the agar plates using a glass L spreader which was sterilized after each use using a flame. The plating of KO transformation used this same process but with the ‘KO rxn’ tube and ‘KO rxn’ agar plate respectively.
The plates were left to sit for about five minutes to ensure that the liquid could soak into the agar. They were then incubated upside down in a 37 °C incubator overnight. It was moved into a 25 °C incubator the next day and the results were observed a week later.
Eight new plates were prepared one week after the CRISPR experiment by heating a bottle of LB agar in 30-second increments until the solution was no longer cloudy. The bottle was left to cool until it could be touched by bare hands and then poured into each plate with a thickness of about three millimeters. Lids were immediately placed on the plates to prevent contamination and the agar was left to solidify.
The ‘Ctrl rxn’ and ‘KO rxn’ plates were taken out of the incubator and observed to verify their success. Successful ‘Ctrl rxn’ colonies appear blue while successful ‘KO rxn’ colonies appear white. Colonies were spread evenly using inoculation loops onto the new agar plates, making four new plates of ‘KO rxn’ and four new plates of ‘Ctrl rxn.’ Each agar plate was labelled ‘KO rxn’ or ‘Ctrl rxn’ for the bacteria it contained. After being spread, the plates were flipped upside down and left in a 25 °C incubator for one week.
The antibiotics used in this study were added in a disk diffusion method. The antibiotics included ampicillin, tetracycline, chloramphenicol, and streptomycin. Each antibiotic came in the form of magazines. Each antibiotic was assigned two agar plates, one ‘KO rxn’ and one ‘Ctrl rxn.’ Each agar plate was labelled with the assigned antibiotic and received four disks of the antibiotic that were evenly distributed across the plate. The agar plates were left to sit until the disks stuck to the bacteria and agar, and then flipped upside down and returned to the 25 °C incubator. The agar plates were left in the incubator for another week and then taken out for observation of results.
Results
The CRISPR experiment resulted in knock-out colonies appearing white and control colonies appearing blue, as expected (Figure 1). However, the growth of the bacteria colonies was minimal and did not supply enough for the second part of this experiment, prompting the further spreading of bacteria (Figure 2).

Figure 1. CRISPR results. Blue colonies are control, white colonies are successful knock-outs.
Figure 2. Bacteria spread onto new plates.
The results of the disk diffusions were minimal and not clear to the naked eye across all groups as there were no observable zones of inhibition around the discs. The disk diffusion did not appear to affect the bacteria, neither the control nor the knock-out (Figures 3-6).
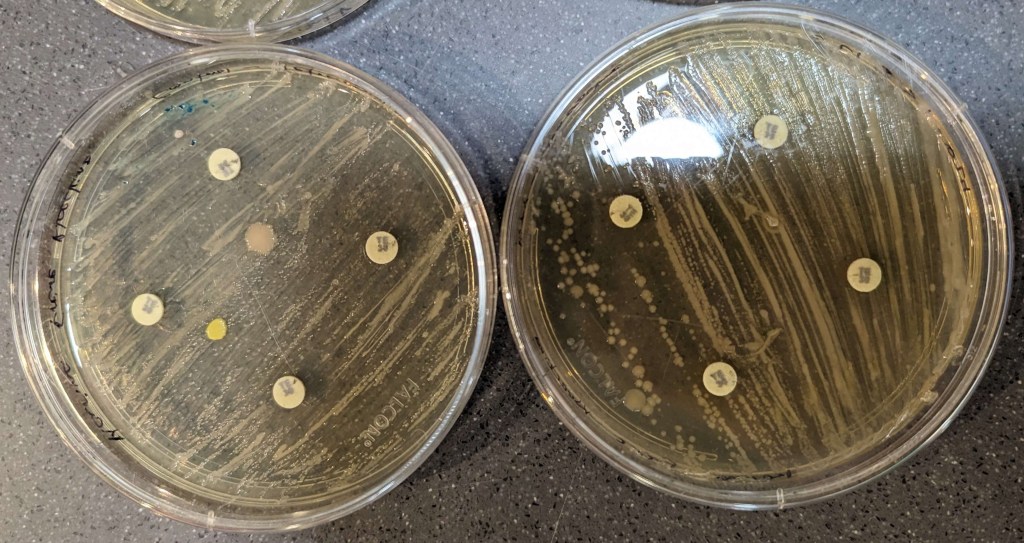
Figure 3. Tetracycline results. Control bacteria is on the left, knock-out bacteria are on the right. The large pink and yellow spots are contamination.
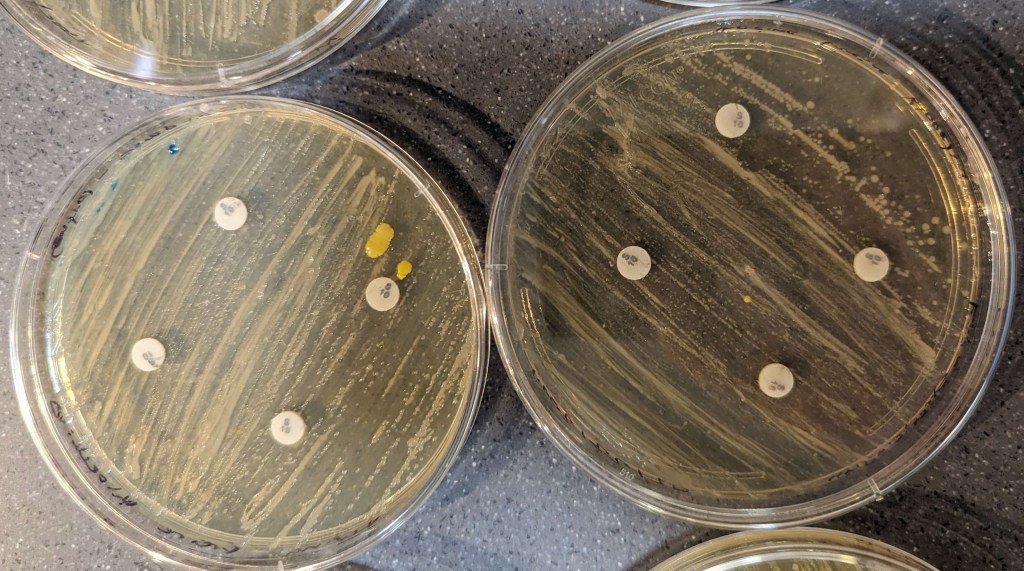
Figure 4. Streptomycin results. Control bacteria is on the left, knock-out bacteria are on the right. The large pink and yellow spots are contamination.
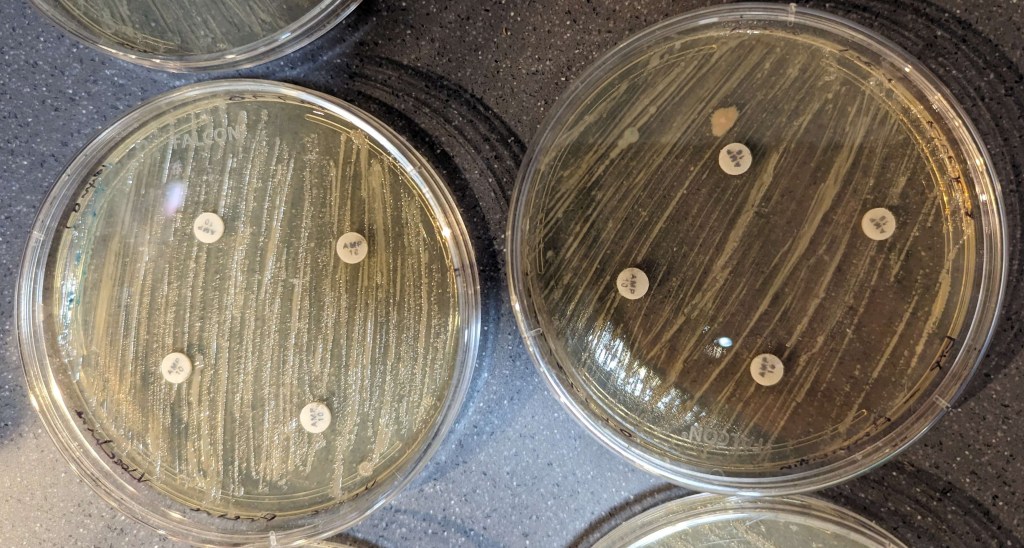
Figure 5. Ampicillin results. Control bacteria is on the left, knock-out bacteria are on the right. The large pink and yellow spots are contamination.

Figure 6. Chloramphenicol results. Control bacteria is on the left, knock-out bacteria are on the right. The large pink and yellow spots are contamination.
Discussion
The results of this study suggest that the antibiotic disk diffusion method did not have discernible effects, proven by the lack of clear inhibition zones for bacterial strains tested (figure 3-6). With no clear difference between the experimental and control assays, the CRISPR experiment likely did not harbor any large off-target effects that would impact the E. coli antibiotic resistance. Instead, the lack of observable effects is most likely attributed to the time at which the disks were added for the disk diffusion method. In the standard assay, disks are added immediately after the spreading of bacteria, (Young, 2021), however, in this case the disks were added a week after spreading. A delayed addition of the antibiotics could lead to changes in bacterial growth phases, and potentially reduce their susceptibility to antibiotics, affecting the formation of inhibition zones. As the error in the method presumably had a large impact on the results, the data should be considered inconclusive, requiring that this experiment be repeated for better results.
Sources of Error and Limitations
Several limitations could have detrimentally impacted the results of this study. First, although the manufacturer’s manual was followed very closely for the miniPCR “Knockout! A CRISPR/Cas Gene Targeting Lab,” there were some time restraints during the experiment that may have led to less optimal results. Also, the minimal colony growth on the original plates (Figure 1), potentially due to an error during the preparation where some of the bacteria spilled, may have posed as a challenge in accurately assessing the antibiotic resistance. Moreover, the new plates that the bacteria were spread on (Figure 2) did show some contamination, which may have impacted the results. Additionally, with this study focusing on a limited set of antibiotics, it overlooked the potential of testing alternative antimicrobial agents and any combination of therapies and their synergistic effects.
In the spirit of these limitations, future studies could explore a variety of topics. With the opportunity to repeat this study, measures would be taken to correct the experimental errors that occurred, hopefully resulting in more conclusive data. Expanding on the antibiotics and antimicrobial agents tested could also offer a more detailed analysis. Another option, using CRISPR to target specific areas of the bacteria that directly relate to antibiotic resistance could provide more in-depth insights into bacterial vulnerabilities and CRISPR’s effects and potential. Similarly, targeting a gene that is naturally present in the bacteria could potentially offer greater off-target effects, as the lacZ gene removed in this study was introduced in a plasmid to the E. coli, and was not naturally present. Branching into the realm outside of bacteria, using CRISPR on plants and examining their overall fitness after the modification could provide further insight into the CRISPR, especially as its abilities are being expanded for a variety of uses.
References
Alteri, C. J., & Mobley, H. L. (2012). Escherichia coli physiology and metabolism dictates adaptation to diverse host microenvironments. Current Opinion in Microbiology, 15(1), 3–9. https://doi.org/10.1016/j.mib.2011.12.004
Battad-Bernardo, E., McCrindle, S. L., Couperwhite, I., & Neilan, B. A. (2004). Insertion of an E. coli lacZ gene in Acetobacter xylinusfor the production of cellulose in whey. FEMS Microbiology Letters, 231(2), 253–260. https://doi.org/10.1016/s0378-1097(04)00007-2
Beneditta Suwono, Hammerl, J. A., Eckmanns, T., Merle, R., Eigner, U., Lümen, M., Lauter, S., Stock, R., Fenner, I., Boemke, E., & Tenhagen, B.-A. (2021). Comparison of MICs in Escherichia coli isolates from human health surveillance with MICs obtained for the same isolates by broth microdilution. JAC-Antimicrobial Resistance, 3(3). https://doi.org/10.1093/jacamr/dlab145
Guo, C., Ma, X., Gao, F., & Guo, Y. (2023). Off-target effects in CRISPR/Cas9 gene editing. Frontiers in Bioengineering and Biotechnology, 11(1143157). https://doi.org/10.3389/fbioe.2023.1143157
Juers, D. H., Matthews, B. W., & Huber, R. E. (2012). LacZ β-galactosidase: Structure and function of an enzyme of historical and molecular biological importance. Protein Science, 21(12), 1792–1807. https://doi.org/10.1002/pro.2165
Kaczmarek, A., Skowron, K., Budzyńska, A., Grudlewska, K., & Gospodarek-Komkowska, E. (2017). Virulence genes and antimicrobial susceptibility of lactose-negative and lactose-positive strains of Escherichia coli isolated from pregnant women and neonates. Folia Microbiologica, 62(5), 363–371. https://doi.org/10.1007/s12223-017-0506-y
Liu, G., Olsen, J. E., & Thomsen, L. E. (2019). Identification of Genes Essential for Antibiotic-Induced Up-Regulation of Plasmid-Transfer-Genes in Cephalosporin Resistant Escherichia coli. Frontiers in Microbiology, 10. https://doi.org/10.3389/fmicb.2019.02203
Lorenzo, D., Esquerda, M., Palau, F., Cambra, F. J., & Bioética, G. I. en. (2022). Ethics and Genomic Editing Using the Crispr-Cas9 Technique: Challenges and Conflicts. NanoEthics, 16(3), 313–321. https://doi.org/10.1007/s11569-022-00425-y
Poirel, L., Madec, J.-Y., Lupo, A., Schink, A.-K., Kieffer, N., Nordmann, P., & Schwarz, S. (2018). Antimicrobial Resistance in Escherichia coli. Microbiology Spectrum, 6(4), 10.1128/microbiolspec.ARBA0026-2017. https://doi.org/10.1128/microbiolspec.ARBA-0026-2017
Seier, T., Padgett, D. R., Zilberberg, G., Sutera, V. A., Toha, N., & Lovett, S. T. (2011). Insights Into Mutagenesis Using Escherichia coli Chromosomal lacZ Strains That Enable Detection of a Wide Spectrum of Mutational Events. Genetics, 188(2), 247–262. https://doi.org/10.1534/genetics.111.127746
World Health Organisation. (2023). Antimicrobial Resistance. http://Www.who.int. https://www.who.int/health-topics/antimicrobial-resistance#:~:text=Antimicrobials%20%2D%20including%20antibiotics%2C%20antivirals%2C
Wu, K. J. (2020, October 31). Crispr Gene Editing Can Cause Unwanted Changes in Human Embryos, Study Finds. The New York Times. https://www.nytimes.com/2020/10/31/health/crispr-genetics-embryos.html#:~:text=genetics%2Dembryos.html-
Young, L. S. (2021). The minimum inhibitory concentration of antibiotics: Methods, interpretation, clinical relevance. Pathogens, 10(2), 165. https://doi.org/10.3390/pathogens10020165
Zhang, H., Cheng, Q.-X., Liu, A.-M., Zhao, G.-P., & Wang, J. (2017). A Novel and Efficient Method for Bacteria Genome Editing Employing both CRISPR/Cas9 and an Antibiotic Resistance Cassette. Frontiers in Microbiology, 8. https://doi.org/10.3389/fmicb.2017.00812