Beatrice Leahu – Applied Science
Abstract
The human genome is one of humanity’s greatest mysteries. The human population has a nearly identical genomic sequence; however, alleles account for the genotypic and phenotypic differences. Single Nucleotide Polymorphisms (SNPs) are a common form of genetic variation drastically affecting the efficiency of the stress receptor gene FKBP5 in Homo sapiens. This paper aims to produce a proof-of-concept for an allele frequency model using the outcomes of genetic variation in FKBP5. It aims to increase the accuracy of statistical analysis of allele frequencies and provide patients with greater insight into SNPs, making healthcare one step easier. The allele frequency model functions as expected, utilizing an author-generated database that acted as a proxy for FKBP5 due to data accessibility barriers. Various difficulties, including open-source genetic databases and biodata accessibility, were faced. This study highlights the need for open-source genetic databases, as no conclusive DNA sequences with identified SNPs and clinical outcomes were located, posing a major limitation during the scientific process.
Introduction
DNA
The human genome is a pattern of nucleotides with four possible nitrogenous bases: adenine (A), guanine (G), cytosine (C), and thymine (T), approximately three billion nucleotides long. Human DNA is nearly identical except for genetic variation, which encodes for unique factors. Genetics is a vast and complex field with a large percentage of inaccuracy. Dr. Stephen Kingsmore, the CEO of Rady Children’s Institute for Genomic Medicine, stated: “In one-quarter of cases, doctors sequence a patient’s genome only to find a variant with an unknown impact on health” (Johnson, 2023). Certain genes, such as FKBP5, can be studied to understand phenotypic and genotypic differences predisposing a patient to certain diseases.
Relevance of FKBP5
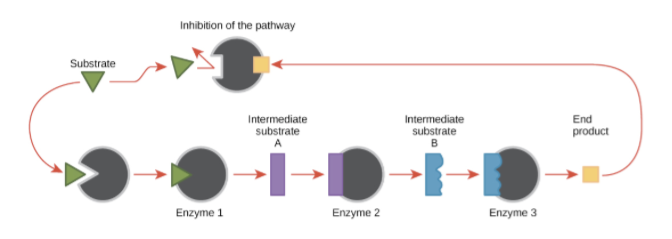
Garth Baughman and his team first discovered FKBP Prolyl Isomerase 5 (FKBP5) in 1995 (Zannas et al., 2016). FKBP5 encodes the information required to produce the FK506-binding protein 51, a cortisol receptor regulator linked directly with mood disorders such as Major Depressive Disorder and Post Traumatic Stress Disorder (Zannas et al., 2016). Cortisol is the body’s core regulating steroid hormone, synthesized from cholesterol. Cortisol has many functions in the human body, for instance, mediating the stress response, regulating metabolism, managing the inflammatory response, and immune function. However, high levels of cortisol have been directly linked to various mental health disorders and have been identified as a factor in nearly every disease (Thau et al., 2023). The protein FK506 co-chaperones and changes the function of other proteins, controlling a negative feedback loop to inhibit the production of cortisol in the adrenal glands. A negative feedback loop occurs in a series of protein production processes (Figure 1). If there is too much of a protein produced, then the entire cycle is terminated. An increase in one protein leads to a decrease in the remaining proteins. This is meant to maintain efficiency and regulate metabolic processes. Thus, FKBP5’s ability to inhibit cortisol production has been correlated with an individual’s ability to develop or manage mood disorders. Genetic variation in FKBP5 can affect how FK506 proteins are encoded, changing their function in cortisol level maintenance and impacting one’s ability to develop or manage mood disorders.
Figure 1: Negative Feedback Inhibition Diagram (Khan Academy, 2024)
SNPs
Single nucleotide polymorphisms (SNPs) are crucial to the biological outcomes of FKBP5. SNPs are one of the most abundant genetic variations in the human genome. They are a crucial biological marker of genetic variation, with the possibility of being advantageous. SNPs are the change of one alternate base pair in a definitive position of the gene. This can occur due to epigenetics, replication errors, and mutations, among other causes. SNPs commonly occur every 300 base pairs of sequence with a minor allele frequency greater than 1% (Nelson et al., 2004). SNPs are most commonly found in introns, which are the non-coding portions of the gene; however, they play a big role in how the protein functions. SNPs can affect the human body in a multitude of ways, including changing the encoded amino acids in proteins, influencing gene expression, changing messenger ribonucleic acid (mRNA) stability, or nothing, leading to possible diseases or fitness advantages (Shastry, 2009). Understanding SNPs and their associated outcomes allows patients to better understand their own predisposition to certain conditions and may allow them to plan for potential medical needs.
Allele Frequency Model
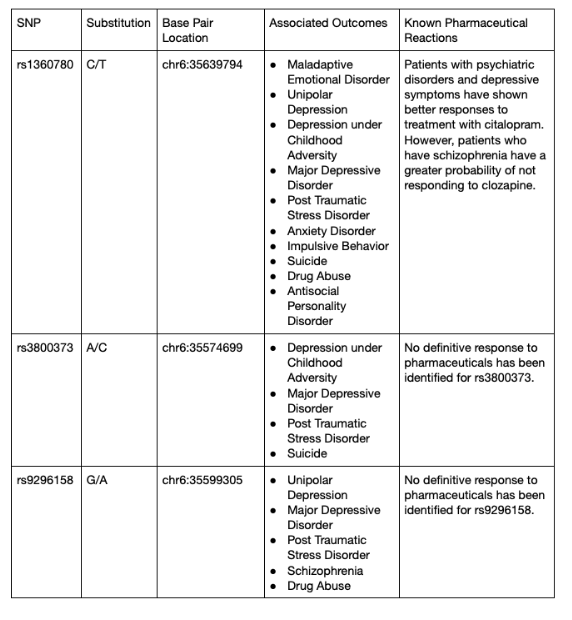
However, scientific papers to understand the complexity of SNPs and genetic variation are dense and inaccessible to the public. This creates a knowledge barrier, preventing patients from educating themselves about possible symptoms and viable courses of action. Accessible biotechnology to educate patients and the public on genetic variation is limited. Most software is designed for medical professionals to identify genes and devise possible genetic therapy solutions. In the last decade, deep mutational scanning has been introduced. Deep mutational scanning processes large-scale datasets that can reveal intrinsic protein properties and protein behavior within cells due to human genetic variation (Fowler et al., 2014). Allele frequency models are programs that calculate the prevalence of an allele in a population to track the appearance of alleles to further draw conclusions in studies regarding epigenetics or heredity. This project aims to develop a program that medical professionals can use to update and calculate allele frequency while simultaneously educating the public, using variation in FKBP5 as an example. To complete this proof-of-concept in a scalable manner, three SNPs were studied by identifying the location of the allele change in the genome, the nucleotide replacement, the possible diseases the SNP can predispose you to, and if there any known pharmaceutical reactions to the SNP (Table 1) (Malekpour et al., 2023).
Table 1: Information Released to Program Users for 3 Different SNPs Studied (Malekpour et al., 2023)
Patients would have to be genetically sequenced prior to using the program. This tool would be available to patients and recommended by medical professionals. The allele frequency model can serve as an add-on to deep mutational scanning, providing more statistical population analysis and increasing accessibility.
Materials and Methods
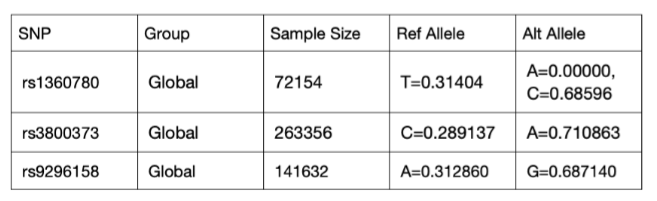
A literature review was conducted to source FKBP5 sequences for analysis. However, due to difficulties in accessing suitable genetic sequences, as highlighted in the discussion, a proxy database was generated. Randomly generated sequences with 50 nucleotides were created by using a random DNA sequence generator from Dr. Maduro’s lab at the University of California Riverside, with 0.50 GC and 0.50 AT base pairs (Random DNA Generator). This sequence was duplicated 100 times in separate rows. These sequences were designed to represent the intron in which FKBP5 SNPs are located. To successfully produce a proof-of-concept, SNP locations were randomly chosen within the span of 50 nucleotides since human SNP locations are thousands of nucleotides apart. Thus, the three SNPs of FKBP5 studied in this experiment—rs1360780, rs3800373, and rs9296158—are placed in locations 5, 25, and 42, respectively. The database was designed so different variants were presented in proportional frequencies following current allele frequency statistics from the National Library of Medicine (Table 2).
Table 2: ALFA Allele Frequency Statistics (rs1360780 RefSNP Report, dbSNP- NCBI, 2022), (rs3800373 RefSNP Report, dbSNP-NCBI, 2022), (rs9296158 RefSNP Report, dbSNP-NCBI, 2022)
The allele frequency program manipulated an array, which tested and transformed the data to be used for various functions. A database of 100 sequences was imported as a comma-separated value (CSV) file.
The program was written in Python through Visual Studio Code by Microsoft. Work packages used include Pylance by Microsoft, Python Debugger by Microsoft, and Rainbow CSV by mechatroner. The model identifies and counts the sequences that exhibit rs1360780, rs3800373, or rs9296158 allele substitutions. After establishing the baseline population data from the sequences provided, the model asked the user to select which SNP they exhibited. Users could select from the current SNP options for FKBP5. One option could be selected or all three. The program then calculated the percent of patients that exhibited the selected genetic variation and printed the percentage of the population that also exhibited the SNP, a brief description of possible symptoms, and a medical advice disclaimer reminding the patient to rely on a medical professional for personalized results. Users were then given the option to participate by inputting their DNA sequence into the database as well. If yes was selected, the sequence was imputed into the original database, creating a better representation of the population over time and gradually improving the accuracy of the program.
Results
Overall, the model fulfills its purpose and runs as expected as a proof of concept. When a user input was submitted, the program output the relevant information for the SNP selected. The correct proportions of each SNP entered in the database, and information was output. An example of the output can be found in Figure 2. If the user agreed to input their own DNA sequences, the program was able to successfully update the population data accurately. The program is self-sustaining, leveraging crowdsourcing to improve the accuracy of the allele frequency model with every use.
Figure 2: Sample User Output
Discussion
The project was successful in managing an array and detecting the relevant information required for a user’s request. The model can be scaled up as necessary to include more SNPs from various genes aside from FKBP5. Various features could be added, such as utilizing AI to research and summarize academic studies to inform users. A future iteration of this model could allow the user to input a genetic sequence and identify which variant they exhibit rather than relying on medical professionals to tell a patient, in addition to information about outcomes. The proof-of-concept was successful and provides a unique approach to accessible knowledge that should be implemented in healthcare. It would be most beneficial if a real genetic database with the correct length of identified introns were accessed, as this form of testing was unattainable. A randomly generated database was utilized to proceed, which does not accurately represent the diversity of human sequences.
Throughout this study, the scope has changed multiple times to account for the various barriers faced. Genes with studied SNPs are often billions of base pairs long with complicated intron and exon patterns. Published sequences are demanding to decipher due to the lack of metadata and clear biological markers. The use of various genetic databases, including GenBank from the National Library of Biomedical Information, was attempted. It is the largest genetic sequence database in North America, with an annotated collection of all publicly available DNA sequences that rely on public submissions of DNA and mRNA sequences from researchers and doctors.
Following hours of research, a usable DNA sequence with definitive SNPs was not located. Firstly, FKBP5 in Homo sapiens was considered. However, only mRNA sequences were available. SNPs for FKBP5 are located in the introns of the gene, thus would not be located in the mRNA sequence. Next, finding FKBP5 in Mus musculus and FKBP5 in Rattus rattus was attempted. Once again, there were no available genetic sequences. Various other genes were then considered, including Insulin (INS) and the Sex-Determining Region Y (SRY). These genes are more researched and less complex; however, there are few to no studied SNPs due to the low degree of genetic variation. Furthermore, genes were researched in Caenorhabditis elegans. Again, the search was inconclusive. Lastly, the prokaryotes Escherichia coli and Salmonella enterica were researched. Although prokaryote sequences are more accessible, similar functions in SNPs have not been identified between eukaryotes and prokaryotes. Biodata from more obscure databases was investigated, including Genecard and the Leiden Open Variation Database (LOVD).
No conclusive DNA sequences, SNPs, and clinical outcomes were located, posing a major limitation during the scientific process. Previous studies sequenced the DNA required to complete their research independently, with funding. However, at the secondary school level, the funding and resources required for such a large-scale project are limited, requiring the creation of a proxy database. Therefore, in the future, it would be beneficial to complete this experiment without relying on open-source databases. The experiment should be repeated with a more definitive database by obtaining genes from research facilities or patients.
Various barriers during the completion of this paper highlighted the inaccessibility of scientific data. Journals and studies restrict the availability of raw data for various reasons, such as privacy, ethical standards, or data mining for future publications (Miyakawa, 2020). The lack of crucial data for experiment replication poses a barrier to scientific innovation. The scientific community is facing a reproducibility crisis, where only a small percentage of papers are reproducible. The “[a]bsence of raw data means the absence of science” (Miyakawa, 2020). It has become common for data to be falsified or fabricated since revealing raw data could discredit a scientist’s work. Nonetheless, this is a growing issue that must be addressed to prevent misinformation and advance scientific innovation.
As innovation in genetics continues to grow, it would be beneficial to create programs to assist medical professionals with understanding the correlation between specific genetic SNPs and their resulting proteins. Genetic sequences and crucial information concerning the human genome are concealed in dense literature, making it difficult for patients to educate themselves. This highlights the need for accessibility to scientific research around the world. Scientists are unable to access generalized sequencing data with clear identification.
References
Denny, W. B., Valentine, D. L., Reynolds, P. D., Smith, D. F., & Scammell, J. G. (2000). Squirrel monkey immunophilin FKBP51 is a potent inhibitor of glucocorticoid receptor binding. Endocrinology, 141(11), 4107–4113. https://doi.org/10.1210/endo.141.11.7785
Enzyme regulation (article) | Khan Academy. (n.d.). Khan Academy. https://www.khanacademy.org/science/ap-biology/cellular-energetics/environental-impacts-on-enzyme-function/a/enzyme-regulation
Höhne, N., Poidinger, M., Merz, F., Pfister, H., Brückl, T., Zimmermann, P., Uhr, M., Holsboer, F., & Ising, M. (2014). FKBP5 genotype-dependent DNA methylation and mRNA regulation after psychosocial stress in remitted depression and healthy controls. The International Journal of Neuropsychopharmacology, 18(4), pyu087. https://doi.org/10.1093/ijnp/pyu087
Fowler, D. M., & Fields, S. (2014). Deep mutational scanning: a new style of protein science. Nature methods, 11(8), 801–807. https://doi.org/10.1038/nmeth.3027
Johnson, M. (2023, June 5). New AI tool searches genetic haystacks to find disease-causing variants. Washington Post. https://www.washingtonpost.com/science/2023/06/01/primate-ai-genome-variants/
Malekpour, M., Shekouh, D., Safavinia, M. E., Shiralipour, S., Jalouli, M., Mortezanejad, S., Azarpira, N., & Ebrahimi, N. D. (2023). Role of FKBP5 and its genetic mutations in stress-induced psychiatric disorders: an opportunity for drug discovery. Frontiers in psychiatry, 14, 1182345. https://doi.org/10.3389/fpsyt.2023.1182345
Miyakawa, T. (2020). No raw data, no science: another possible source of the reproducibility crisis. Molecular Brain, 13(1). https://doi.org/10.1186/s13041-020-0552-2
Nelson, M. R., Marnellos, G., Kammerer, S., Hoyal, C. R., Shi, M. M., Cantor, C. R., & Braun, A. (2004). Large-scale validation of single nucleotide polymorphisms in gene regions. Genome research, 14(8), 1664–1668. https://doi.org/10.1101/gr.2421604
Random DNA Generator. http://www.faculty.ucr.edu/~mmaduro/random.htm.
rs1360780 RefSNP Report – dbSNP – NCBI. (2022, September 21). dbSNP. https://www.ncbi.nlm.nih.gov/snp/rs1360780
rs3800373 RefSNP Report – dbSNP – NCBI. (2022, September 21). dbSNP. https://www.ncbi.nlm.nih.gov/snp/rs3800373
rs9296158 RefSNP Report – dbSNP – NCBI. (2022, September 21). dbSNP. https://www.ncbi.nlm.nih.gov/snp/rs9296158
Shastry, B. S. (2009). SNPs: impact on gene function and phenotype. Methods in molecular biology (Clifton, N.J.), 578, 3–22. https://doi.org/10.1007/978-1-60327-411-1_1
Tang, J., Leunissen, J. A., Voorrips, R. E., van der Linden, C. G., & Vosman, B. (2008). HaploSNPer: a web-based allele and SNP detection tool. BMC genetics, 9, 23. https://doi.org/10.1186/1471-2156-9-23
Thau, L., Gandhi, J., & Sharma, S. (2023, August 28). Physiology, cortisol. StatPearls – NCBI Bookshelf. https://www.ncbi.nlm.nih.gov/books/NBK538239/
Zannas, A. S., Wiechmann, T., Gassen, N. C., & Binder, E. B. (2016). Gene-Stress-Epigenetic Regulation of FKBP5: Clinical and Translational Implications. Neuropsychopharmacology : official publication of the American College of Neuropsychopharmacology, 41(1), 261–274. https://doi.org/10.1038/npp.2015.235